What is PCR and how does it work?
Regions of DNA that are chosen for amplification are both anonymous and very specific to the individual. Some of our DNA contains relatively short sequences that are repeated over and over in a row. The number of these repeats usually vary from individual to individual and the repetition patterns are inherited in a Mendelian fashion. There are two types of repeating sequences: variable number tandem repeats (VNTR's) and short tandem repeats (STR's). The small amounts of DNA need to be amplified before they can be analyzed and they can be amplified by PCR.
The PCR procedure was developed by Kary Mullis in 1983, and he earned the Nobel Prize for chemistry in 1993. This Nobel Prize was shared by Mullis with Michael Smith who was from UBC and Smith developed a technique for changing one base at a time in DNA which is called site-directed mutagenesis. PCR is used in gene mapping, cloning, DNA sequencing, gene detection and DNA profiling. It is routinely used as a medical diagnostic tool, in criminal investigations and in courts of law to identify defenders on the molecular level.
The PCR procedure was developed by Kary Mullis in 1983, and he earned the Nobel Prize for chemistry in 1993. This Nobel Prize was shared by Mullis with Michael Smith who was from UBC and Smith developed a technique for changing one base at a time in DNA which is called site-directed mutagenesis. PCR is used in gene mapping, cloning, DNA sequencing, gene detection and DNA profiling. It is routinely used as a medical diagnostic tool, in criminal investigations and in courts of law to identify defenders on the molecular level.
|
The DNA sample is amplified by PCR through copying the sample over and over. In order to do PCR you start with a template strand of DNA that was isolated from the crime scene. Primers are then added to the strand, these primers are short oligonucleotides of DNA chosen to bind at the desired place on the DNA (Kluftinger, J. and Plunket, R., 2015).
Basic PCR required four components: a DNA sample with the segment for amplification , a pair of synthetic oligonucleotides primers, deoxynucleoside triphosphates (dNTP's) and DNA polymerase (Nelson et al., 2013). When amplifying VNTR or STR regions the DNA sequences that flank the sequences are identical in almost all humans and therefore they are excellent templates for the primers. Once the primers are annealed then the DNA needs to extended and this is done by Taq polymerase. The reason that a normal polymerase cannot be used in PCR is because it will get denatured when the strands are denatured. Taq polymerase is special is because it was isolated from the bacterial Thermus aquaticus that are heat resistant because they were isolated from a hot springs in Yellowstone National Park, Wyoming (Kluftiner, J. and Plunket, R., 2015). PCR also requires deoxynucleoside triphosphates (dNTP's) and magnesium chloride which is required for the reaction. There are four steps that occur in PCR:
The PCR cycle conditions are 2:00 at 94°C + 35[30 sec at 94°C + 30 sec at 52°C + 1:00 at 72°C] + 10 min at 72°C + infinity at 4°C |
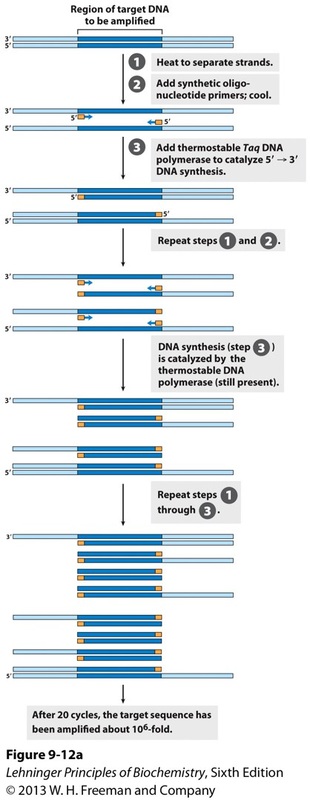
Figure 3. The PCR procedure which has 3 steps. First the DNA strands are separated by heating, the primers are annealed to the separated strands to enable amplification. The new DNA is synthesized by polymerization catalyzed by Taq polymerase. The three steps are repeated for 25 to 30 cycles (Nelson et al., 2013).
|
https://www.youtube.com/watch?v=2KoLnIwoZKU